SPANDx workflow for analysis of haploid next-generation re
$ 14.00 · 5 (263) · In stock


Comparative genomic analysis identifies X-factor (haemin)-independent Haemophilus haemolyticus: a formal re-classification of 'Haemophilus intermedius

Derek SAROVICH, Senior Research Fellow, PhD Microbiology, University of the Sunshine Coast, USC, Faculty of Science, Health, Education and Engineering

VDAP-GUI: a user-friendly pipeline for variant discovery and annotation of raw next-generation sequencing data
My Biosoftware – Bioinformatics Softwares Blog – Page 894 – Supply Bioinformatics Softwares Everyday

Metabolism of ʟ -arabinose converges with virulence regulation to promote enteric pathogen fitness

Comparative genomics and antimicrobial resistance profiling of Elizabethkingia isolates reveals nosocomial transmission and in vitro susceptibility to fluoroquinolones, tetracyclines and trimethoprim-sulfamethoxazole
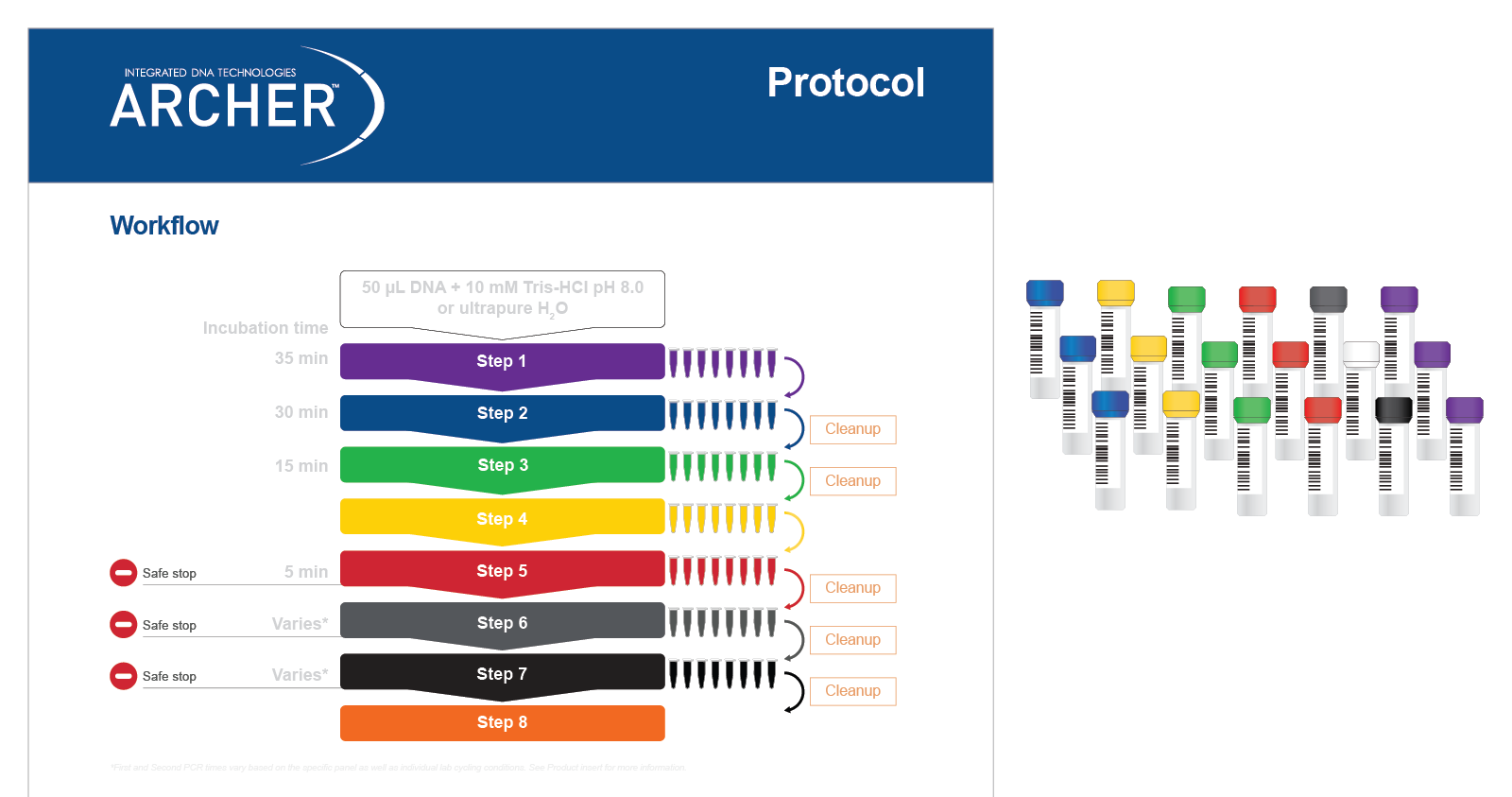
Solid Tumor Research Assays - Archer NGS

PDF] The Genome Analysis Toolkit: a MapReduce framework for analyzing next- generation DNA sequencing data.
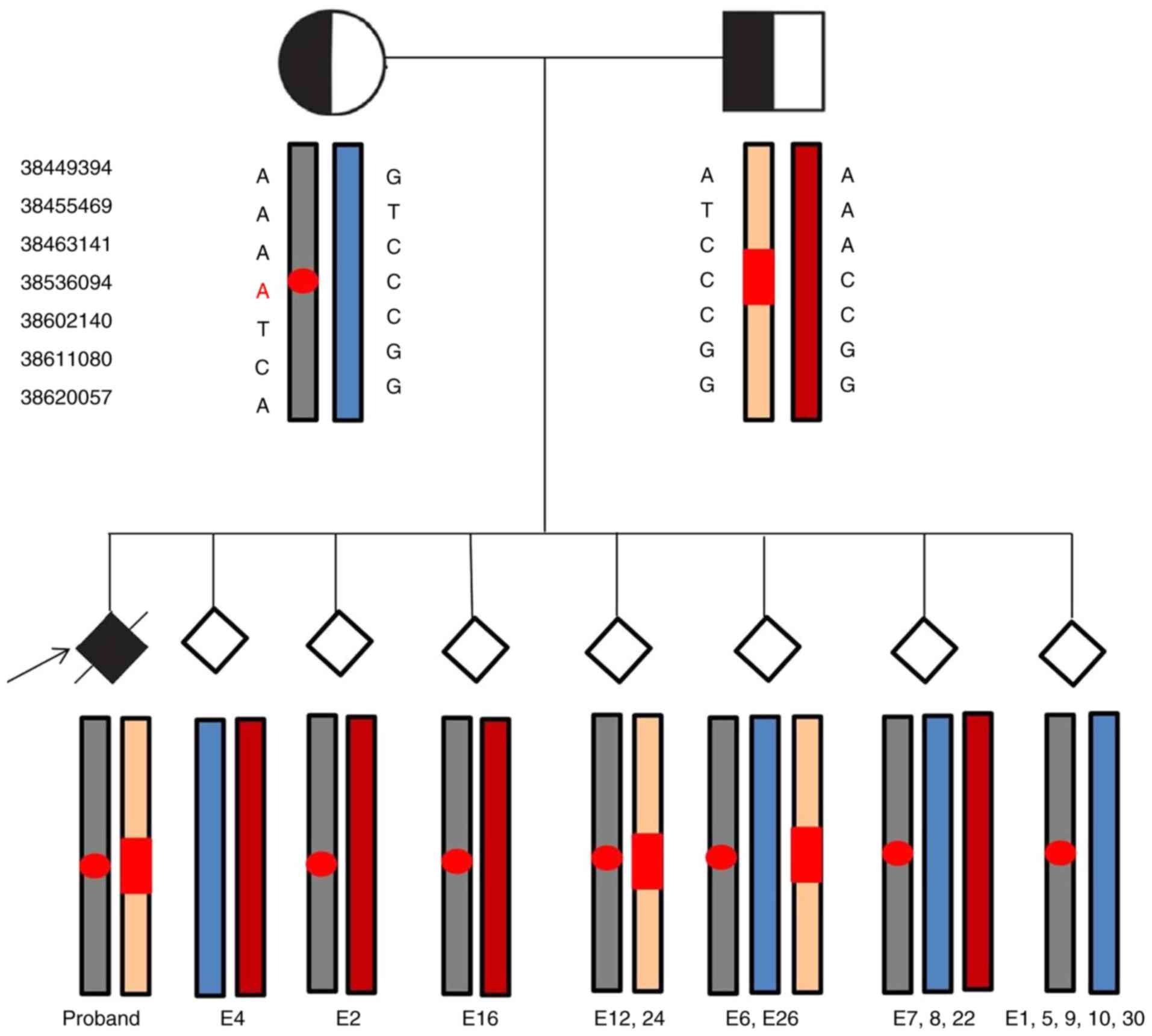
Successful clinical application of pre‑implantation genetic diagnosis for infantile neuroaxonal dystrophy

SPANDx workflow for analysis of haploid next-generation re-sequencing data.
Transformation-associated recombination (TAR) cloning and its applications for gene function; genome architecture and evolution; biotechnology and biomedicine
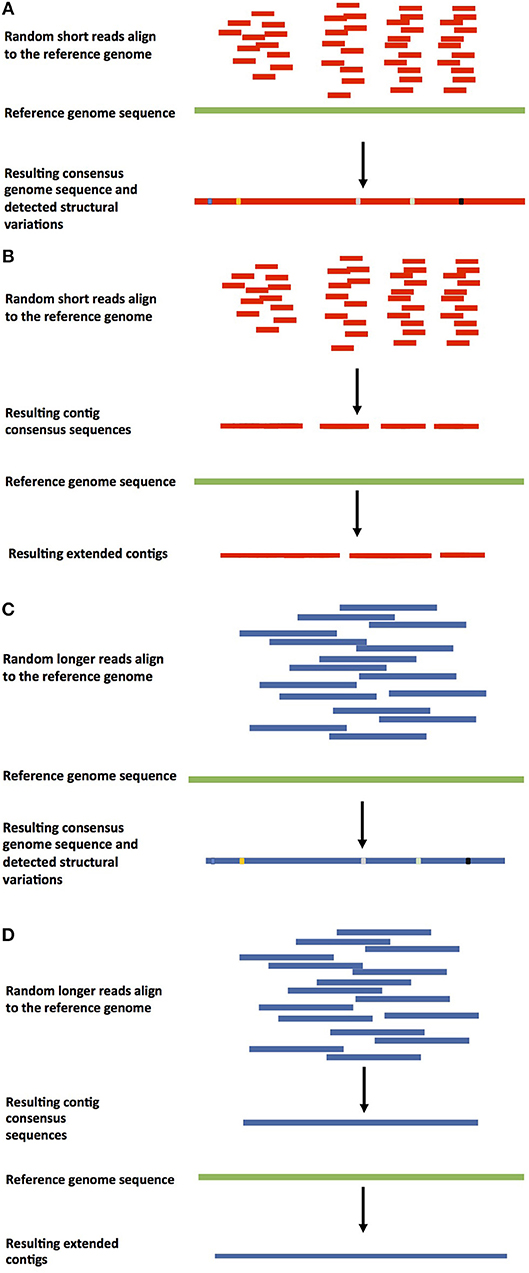
Frontiers Current Strategies of Polyploid Plant Genome Sequence Assembly