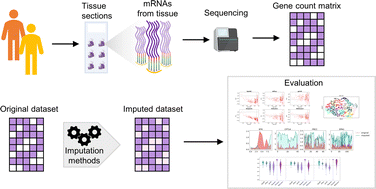
Low-pass sequencing and imputation for evaluating genetic
$ 12.99 · 4.9 (600) · In stock
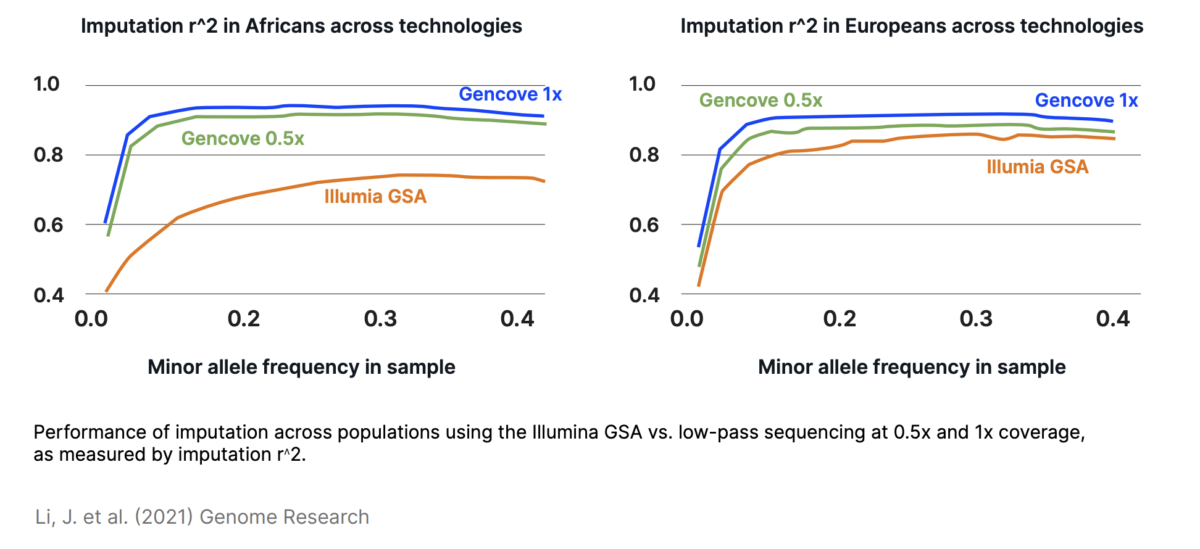
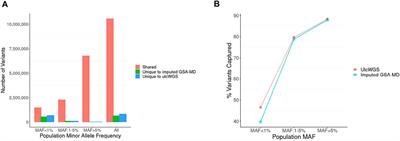
Frontiers Ultra Low-Coverage Whole-Genome Sequencing as an Alternative to Genotyping Arrays in Genome-Wide Association Studies
A comparative performance evaluation of imputation methods in spatially resolved transcriptomics data - Molecular Omics (RSC Publishing)

Quoteworthy: How Swine and Poultry Leaders Drive Genetic Improvement
Low-Pass Whole Genome Sequencing

67 Free DNA Sequence Analysis Tools - Software and Resources
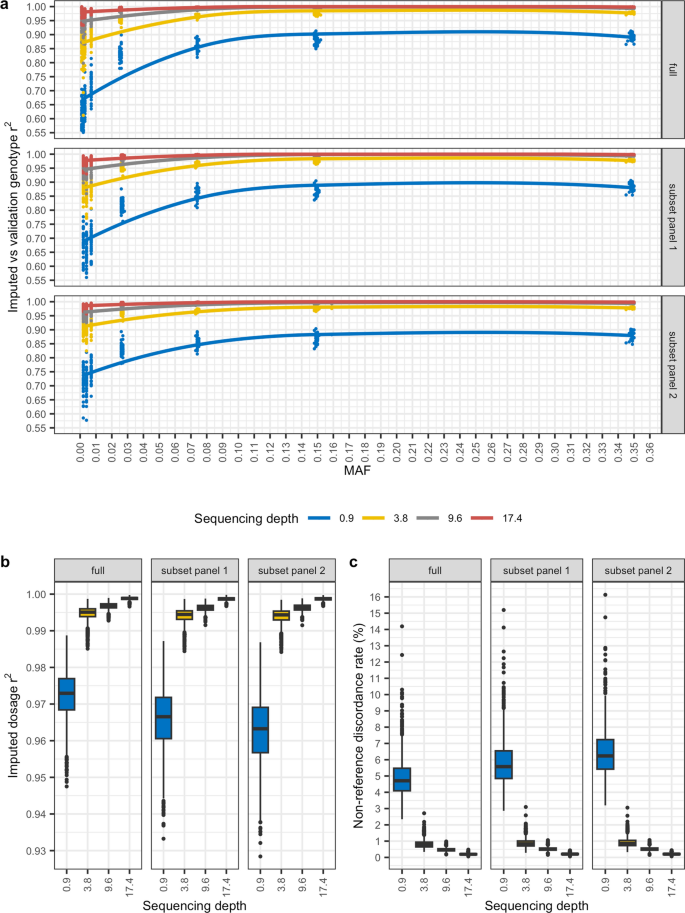
A cautionary tale of low-pass sequencing and imputation with respect to haplotype accuracy, Genetics Selection Evolution

Low-Coverage Whole Genome Sequencing - NCI

Assessment of the performance of different imputation methods for low-coverage sequencing in Holstein cattle - ScienceDirect

Benchmarking algorithms for pathway activity transformation of single-cell RNA-seq data - Computational and Structural Biotechnology Journal
Frontiers Comparison of Genotype Imputation for SNP Array and Low-Coverage Whole-Genome Sequencing Data

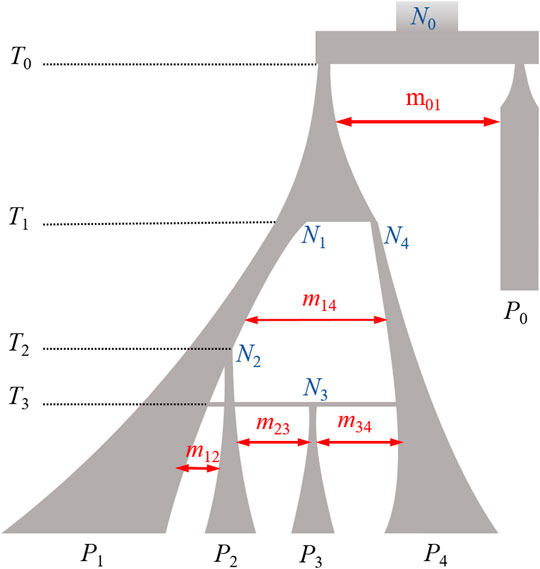
Filtering strategies for reducing imputation errors. a Schematic of
Gencove's new pig haplotype reference panel, by Gencove, The Gencove Blog
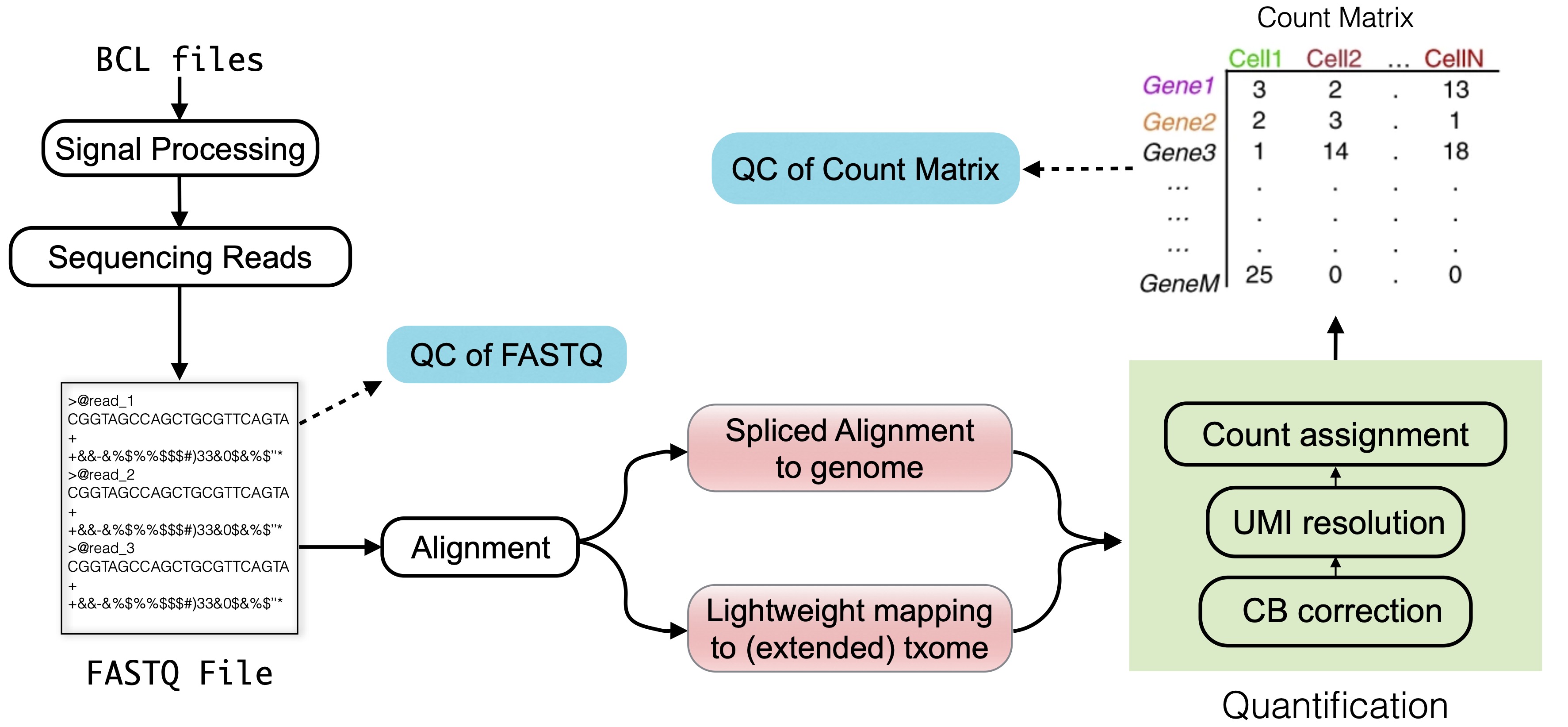
3. Raw data processing — Single-cell best practices